Bifidobacterium animalis subsp. lactis is a Gram-positive lactic acid bacteria commonly found in the healthy human and animal gut. It is prevalent in the infant gut microbiota, ileum, feces, and mucosa, and also found in the intestines of chickens, rabbits, and the gastrointestinal tracts of pigs and dogs [1,2]. This bacterium survives in the gastrointestinal tract, attaches to human epithelial cells in vitro, modulates the fecal microbiota composition, and can prevent gastrointestinal and colonic disorders mediated by microbes [3,4]. These beneficial effects have established B. animalis subsp. lactis as a crucial component in the global industry of functional foods, infant formula, and dietary supplements.
The genus Bifidobacterium belongs to the order Bifidobacteriales, class Actinobacteria, and phylum Actinobacteria. These non-motile anaerobic bacteria are predominantly isolated from the oral and intestinal tracts of mammals, including humans [5]. Comparative studies have shown that B. animalis subsp. lactis has a more streamlined genome compared to other species like Bifidobacterium longum and Bifidobacterium breve, reflecting its specific adaptation to the human gut environment, particularly in infants [6].
This study focuses on the complete genome sequence of B. animalis subsp. lactis KL101 (KL101), isolated from infant feces, highlighting its structural and functional genomic features. By comparing these features with other B. animalis strain such as B. animalis subsp. lactis BB12 (BB12), we aim to provide a deeper understanding of the genomic adaptations that support its probiotic functions and its potential applications in enhancing gut health.
Additionally, this study contributes to animal industry by exploring how the unique genomic traits of KL101 can be leveraged to improve gut health in livestock. Understanding these traits is crucial for developing probiotics that enhance animal health and productivity, thereby supporting more efficient and sustainable animal husbandry practices.
To investigate these features, fecal samples from six-month-old infant were collected using sterile tools and placed in blood liver (BL) liquid medium overlaid with paraffin oil. The samples were stored at 4°C and processed within 24 hours to preserve bacterial viability. The fecal samples were then diluted with sterile phosphate buffer containing 0.05% L-cysteine (5 mM, pH 7.2) and spread on BL-NPNL agar plates for anaerobic incubation at 37°C for 72 hours using an anaerobic incubator. White colonies that were Gram-positive and catalase-negative were isolated. Finally, 16S rRNA analysis confirmed the identity of B. animalis. The identified strain was designated as B. animalis subsp lactis KL101.
KL101 was cultured anaerobically in glucose BL broth at 37°C for 24 hours. First, genomic DNA extraction was conducted using the Exgene Cell SV mini kit (GeneAll), following the manufacturer’s instructions. Next, the complete genome of KL101 was sequenced at Theragen Bio using the Illumina NovaSeq 6000 platform with the TruSeq Nano DNA Sample Prep Kit 150PE (Illumina), generating short reads. Sequence assembly was then performed with fastp (version 0.20.0), Unicycler (version 0.5.0), Pilon (version 1.24), and QUAST (version 5.20). Afterward, the assembled sequences were assessed using BUSCO (version v5.5.0_cv1). Chromosome contig annotation was conducted using Prokka (version 1.14.5-2). Functional annotations for genes and proteins were gathered using eggNOG-mapper (version 2.1.9).
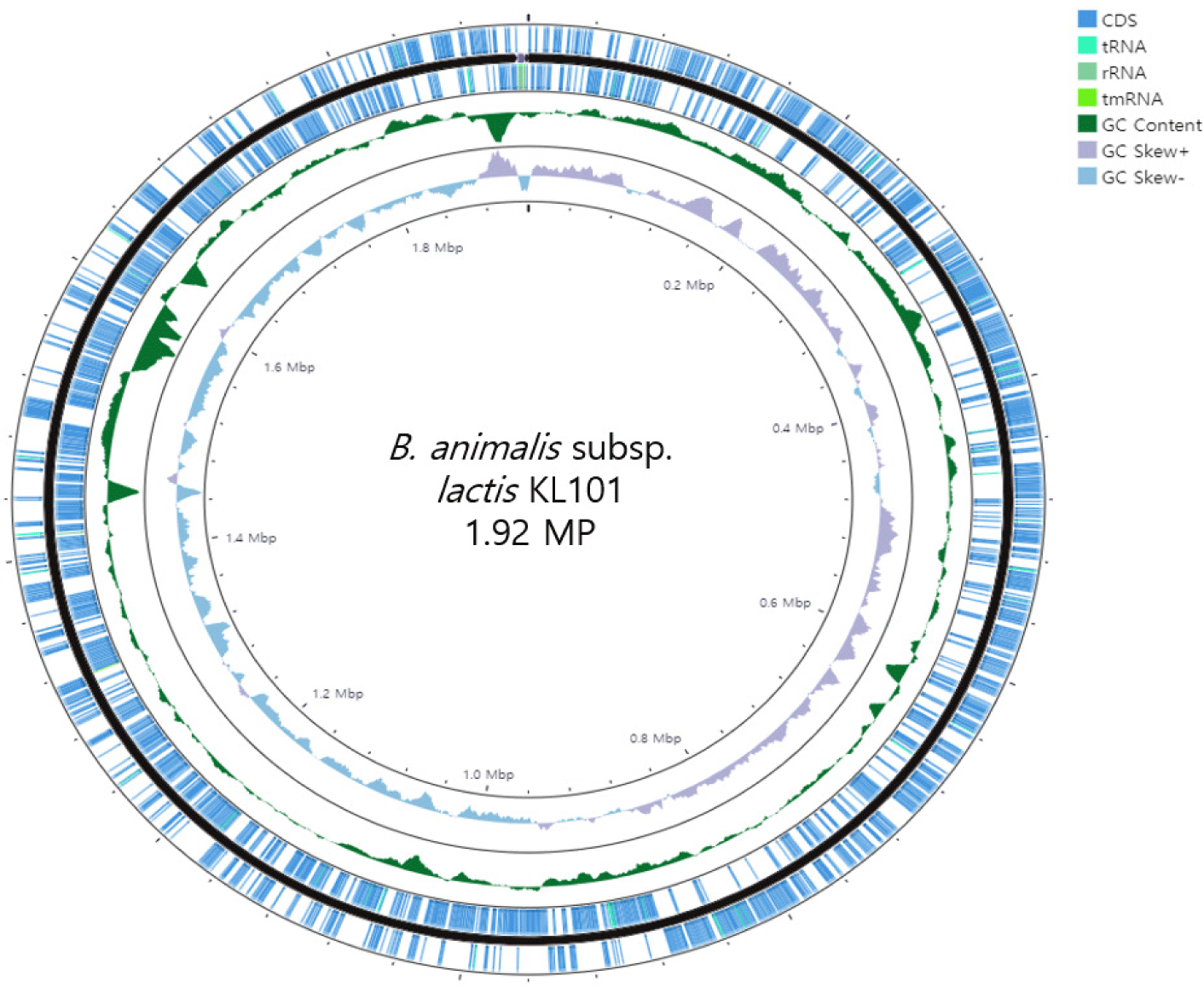
Based on this analysis, the genome of KL101 consists of a single circular chromosome with a total size of approximately 1,919,804 base pairs (bp) (Table 1). This relatively compact genome is characteristic of the Bifidobacterium genus, reflecting a streamlined set of genetic instructions tailored to its ecological niche. The guanine-cytosine (GC) content of the KL101 genome is about 60.4% (Fig. 1), consistent with other bifidobacterial genomes, indicative of its genomic stability and evolutionary adaptations.
| Genomic features | B. animalis subsp. lactis KL101 (Chromosome) |
|---|---|
| Genome size (bp) | 1,919,804 |
| GC content (%) | 60.4 |
| N501) (bp) | 1,912,929 |
| rRNA genes | 3 |
| tRNA genes | 53 |
| tmRNA | 1 |
| CDS | 1,555 |
When compared to other Bifidobacterium species, such as Bifidobacterium longum and Bifidobacterium breve, and KL101 exhibits a smaller genome size. For instance, the genome of B. longum typically spans approximately 2.26 Mb with a GC content of around 60%, while B. breve has a genome size of approximately 2.3 Mb [7,8]. This difference in genome size highlights the evolutionary pressure on KL101 to maintain a streamlined genome that supports its efficient functioning in the infant gut.
KL101 contains 1555 coding sequences (CDSs), 53 tRNAs, and 3 rRNAs (Fig. 2). Various associated genes have also been identified. In the genome of KL101, there are genes encoding bile salt hydrolase (EC 3.5.1.24), which may confer resistance to bile acids. Additionally, the genome also contains thioredoxin system-encoding genes, which are alkyl hydroperoxide reductase C (ahpC), putative peroxiredoxin (bcp), thioredoxin reductase (trxB), peptide methionine sulfoxide reductase (MsrAB), divalent metal cation transporter (MntH), and putative thioredoxin 2 (trxC), suggesting potential utilization as antioxidants in the future [8].
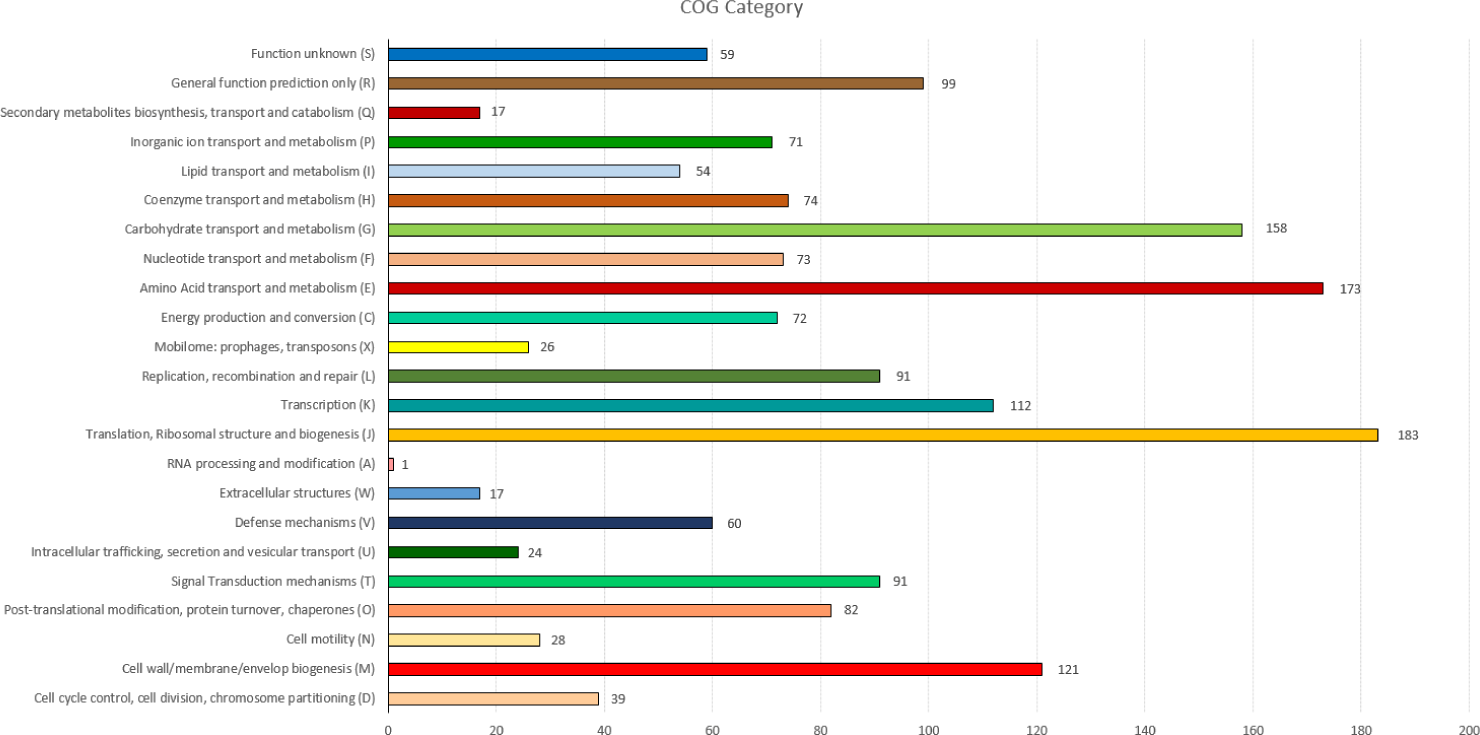
Compared to other bifidobacteria, KL101 shows a unique set of genes that enhance its probiotic functionality. For example, the presence of genes related to the oxidative stress response is more pronounced in KL101 compared to B. longum, which may contribute to its enhanced survival in the harsh conditions of the gastrointestinal tract [9]. Furthermore, universal CRISPR genes cas1 and cas2, which are crucial for adaptive immunity and genomic stability, have been identified in KL101.
A notable feature of the KL101 genome is its extensive array of genes dedicated to carbohydrate metabolism. This includes a variety of glycosyl hydrolases such as β-galactosidases (EC 3.2.1.23) and β-glucosidases (EC 3.2.1.21), which facilitate the degradation of complex carbohydrates into simpler sugars that can be readily absorbed and utilized by the bacterium [10]. The presence of the phosphoketolase pathway (EC 4.1.2.22) is another key element central to the fermentation of pentose and hexose sugars, producing energy and key metabolic intermediates [11].
When compared to other bifidobacteria, such as B. longum and B. breve, KL101 shows a similar but more specialized profile of carbohydrate-active enzymes. These differences underscore the unique adaptations of KL101 to the infant gut, where milk-derived carbohydrates are a primary nutrient source [12]. For example, B. longum has a broader range of carbohydrate metabolism genes, reflecting its adaptation to a more varied adult diet, whereas KL101’s genome is finely tuned to efficiently metabolize milk oligosaccharides found in the infant diet [13,14].
When comparing the genome of KL101 with that of the well-characterized BB12 strain, several distinct and shared features emerge. BB12 is known for its extensive use in probiotics due to its robust health benefits. In addition, both strains share a high degree of similarity in terms of GC content and general genome organization, indicating a conserved genomic architecture within the subspecies. However, KL101 exhibits unique genomic features that may contribute to its specific adaptations in the infant gut.
Additionally, the genome of KL101 (1.92 Mb) is slightly smaller than that of BB12 (1.96 Mb). This difference, albeit minor, may reflect variations in non-essential genes or strain-specific adaptations. Regarding carbohydrate-active enzymes (CAZymes), KL101 and BB12 both exhibit a rich repertoire, enabling the efficient breakdown of dietary fibers and oligosaccharides. This capability is essential for their role in modulating the gut microbiota and enhancing host health. Moreover, both strains possess genes for bile salt hydrolase and the thioredoxin system, which are crucial for survival in the gastrointestinal tract. However, the exact composition and regulation of these genes may vary, contributing to strain-specific probiotic properties.
The presence of CRISPR-Cas systems in both KL101 and BB12 highlights their ability to defend against phage attacks and maintain genomic stability. The specific array of stress response genes, including those for oxidative stress and heat shock proteins, underscores their resilience in the dynamic gut environment.
Overall, the whole genome sequencing and comparative genomic analysis of KL101 provide valuable insights into its probiotic functionalities and adaptations to the infant gut. The unique genomic traits of KL101 underscore its potential for use in feed supplements and health foods aimed at improving gut health. Further research into these genomic features could facilitate the development of targeted probiotic interventions for both human and animal health.
