Lactiplantibacillus plantarum (L. plantarum) used belongs to the genus Lactobacillus, which exists widely in a variety of environments and has been consumed in the context of fermented foods or used by humans for thousands of years. It is a recognized safe microorganism [1]. L. plantarum has many functions, such as maintaining the balance of intestinal flora, improving immunity and promoting nutrient absorption [2]. Studies have shown that dietary L. plantarum can improve the daily gain weight and daily feed intake of weaned piglets, as well as the feed conversion rate [3]. L. plantarum can also regulate the intestinal flora of broilers and accelerate the maturation of broilers [4]. According to previous studies [5], L. plantarum ST isolated from De’ang pickled tea, a traditional food consumed by the De’ ang nationality of Yunnan, China. The probiotic characteristics of this strain were evaluated from several aspects. It was found that L. plantarum ST had good tolerance to gastric and intestinal fluid, strong adhesion and automatic aggregation ability in the intestinal tract, and exhibited antagonistic activity against Salmonella typhimurium and Escherichia coli. Therefore, it is considered that this strain has good probiotic characteristics. At the same time, our previous study has shown that dietary replacement of fermented soy bean hull and canola seed cake with A. niger and L. plantarum ST increased body weight and daily weight gain of 56-day-age Chahua broiler, a local chicken breed in Yunnan of China, compared to that prepared with Aspergillus niger alone (unpublished data). Therefore, L. plantarum ST shows good potential probiotic properties and it maybe can used as a probiotics in animal breeding.
Based on previous studies, the whole genome of L. plantarum ST was analyzed in this paper. L. plantarum ST was cultured in MRS medium (Guangzhou Huankai Microbial Technology, Guangzhou, China) at 37°C for 24 hours. Genomic DNA was extracted from the collected samples using the Wizard® Genomic DNA Purification Kit. PacBio SMRT sequencing process was used to establish a 10-kb library of extracted genomic DNA, after which the whole genome was sequenced using PacBio RS II (Pacific Biosciences, Menlo Park, CA, USA) sequencing platform [6]. L. plantarum ST sequence quality control and strain genome assembly were performed by RS_HGAP_Assembly.3 software in SMRT® Portal (V2.7). Circlator (V1.5.5) [7] was used to cyclize the assembled genome. Upload the assembled L. plantarum ST nucleic acid sequence file to EggNOG database (http://eggnog5.embl.de/) for genome function annotation. Upload the whole genome sequence of L. plantarum ST to CGview server (http://stothard.afns.ualberta.ca/cgview_server/) to complete the genome circle map.
Data evaluation and sequence assembly were conducted for the sequenced sample L. plantarum ST to obtain the genome assembly results (Table 1), and the assembled sequences were spliced into a genome circle map (Fig. 1). The results showed that the L. plantarum ST genome had 1 circular chromosome and 7 plasmids. The whole genome length was 3320817 bp, in which the size of circular chromosome was 3058984 bp and guanine + cytosine (G ± C) content (%) was 44.76%, which contained 2945 protein-coding sequences (CDS). Clusters of Orthologous Groups (COG) annotation results showed that these genes were clustered in 22 functional categories of protein (Fig. 2), among which the number of genes involved in carbohydrate transport and metabolism was the largest, suggesting that this strain has strong carbohydrate metabolism ability. The comparative genomics study of 114 strains of L. plantarum conducted by Mao et al. [8] demonstrated that the core functions of these strains were concentrated on carbohydrate metabolism and amino acid metabolism, which provide molecular support for strains to metabolize multiple sugars. Moreover, the ability of L. plantarum strains to utilize carbohydrates has been correlated with the carbohydrate metabolism-related genes. The COG annotation findings of 126 strains of L. plantarum in another study [9] revealed that the number of coding genes involved in carbohydrate metabolism was the largest, which was consistent with the COG annotation results in this study. Because carbohydrates are the main energy source to support growth and development of microorganisms, lactic acid bacteria can adapt to the environment specifically through obtaining carbohydrates from environment and/or removing unnecessary carbohydrate genes.
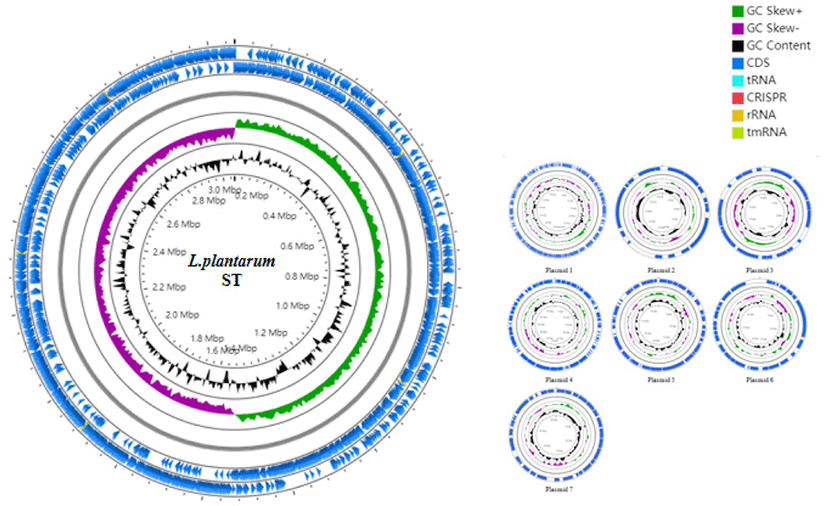
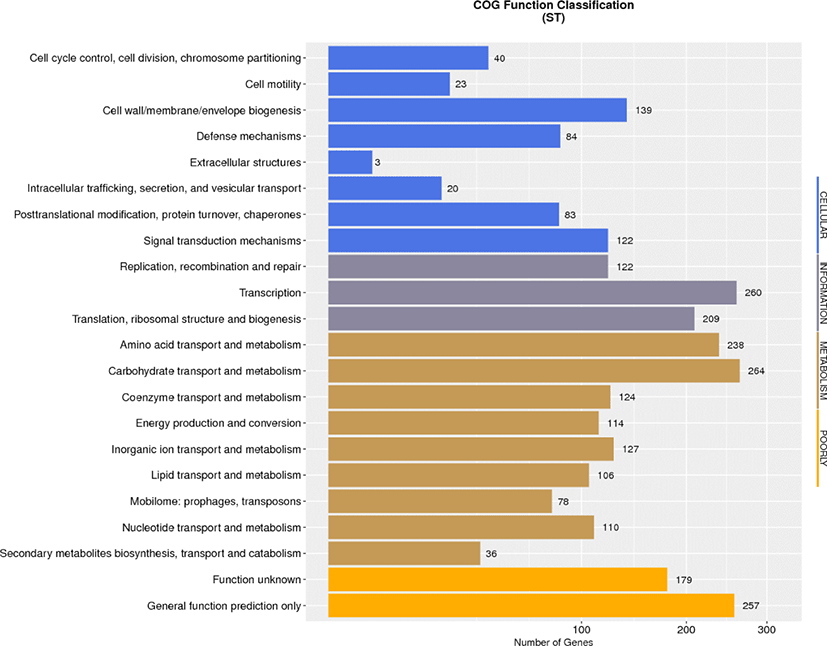
In addition, the genome of ST strain contained 139 genes annotated with cell wall, cell membrane and extracellular membrane biosynthesis, which can provide evidence that this strain has strong biofilm formation ability, and in turn facilitate resistance against external unfavorable factors. Eighty-four annotated genes were identified in the defense mechanism, indicating that the strain could resist the digestive tract environment and provide necessary conditions for stable colonization in intestinal epithelial cells. The defense mechanism also contained four functional sequences synthesizing ABC antibacterial peptide transport system, which is recognized as a necessary factor for antibacterial peptide transport and secretion, and ensure the antibacterial characteristics of the strain. The genomic information of L. plantarum ST provides scientific basis for its future application in animal breeding.
NUCLEOTIDE SEQUENCE ACCESSION NUMBER
L. plantarum ST complete genome sequence has been submitted to GenBank database (https://www.ncbi.nlm.nih.gov/),and deposited under the BioProjecct PRJNA792432.

