INTRODUCTION
Bacteriocin production among lactic acid bacteria is currently a rapidly expanding field of research, which has been stimulated to a large extent by issues regarding the safety of using live probiotics and has, thus, driven a shift toward the development and elaboration of postbiotics [1]. Postbiotics are currently defined as bioactive compounds that are not probiotic, prebiotic, or paraprobiotic, and can include soluble factors, cell compounds, and metabolic products and/or by-products. Compared with live probiotics, some of the advantages of using postbiotics, including bacteriocins, include safer administration, longer shelf-life, and lower probability of the acquisition and spread of resistance genes and other virulence factors [2]. However, although bacteriocin production is considered a favorable trait for potential probiotic strains, the same may not hold true for pathogenic microbes.
In order to successfully colonize a host and establish a suitable niche, pathogens require diverse mechanisms that would enable them to evade possible immune responses, replicate, and ensure disease transmission. Clostridium perfringens, a Gram-positive bacterium, is a known causative agent for several disorders in humans (clostridial myonecrosis and food poisoning) and animals (necrotic enteritis in poultry) [3]. The pathogenicity of C. perfringens is associated with the production of major toxins (α-toxin, β-toxin, ι-toxin, and ε-toxin), subsidiary toxins (collagenase, enterotoxin, and perfringolysin O), and antimicrobial peptide resistance genes (e.g., tetracycline resistance efflux protein and specific aminoglycoside resistance genes) [4]. Here, based on genetic analysis, we report evidence of bacteriocin production by a C. perfringens isolate, provisionally designated as strain B20. Bacteriocin production by a pathogen may enhance its pathogenicity through the same mechanisms employed by probiotic strains to becomes more beneficial; specifically, through direct inhibition of competing strains and niche establishment, as well as through modulating the local microbiota [2]. Thus, it is important to deduce how pathogens acquire genes for bacteriocin production and thereby gain a better understanding of how these traits may affect their pathogenicity.
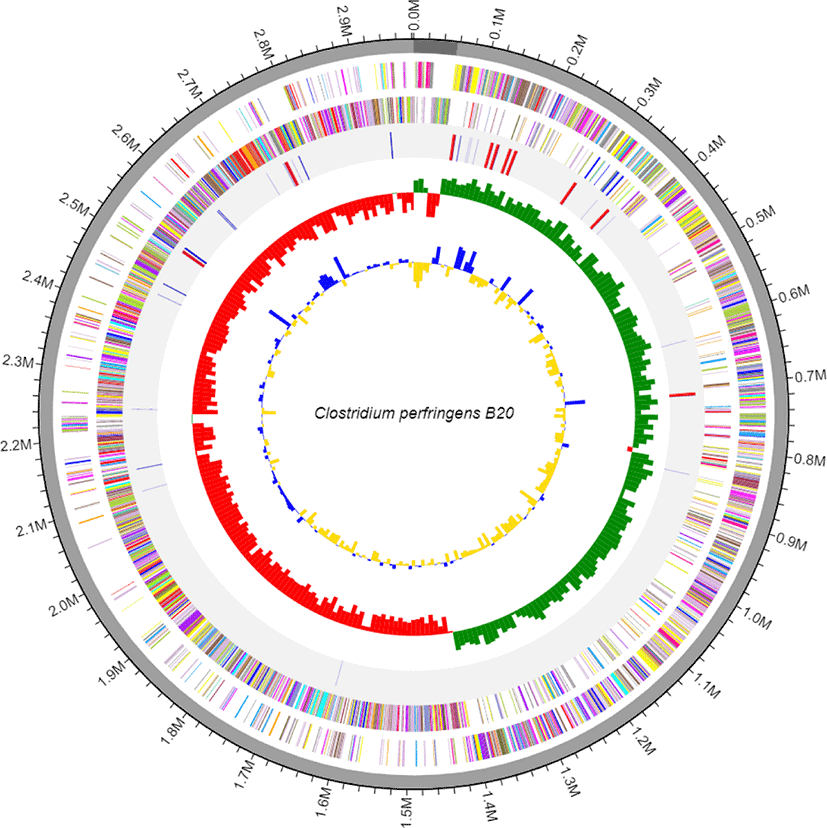
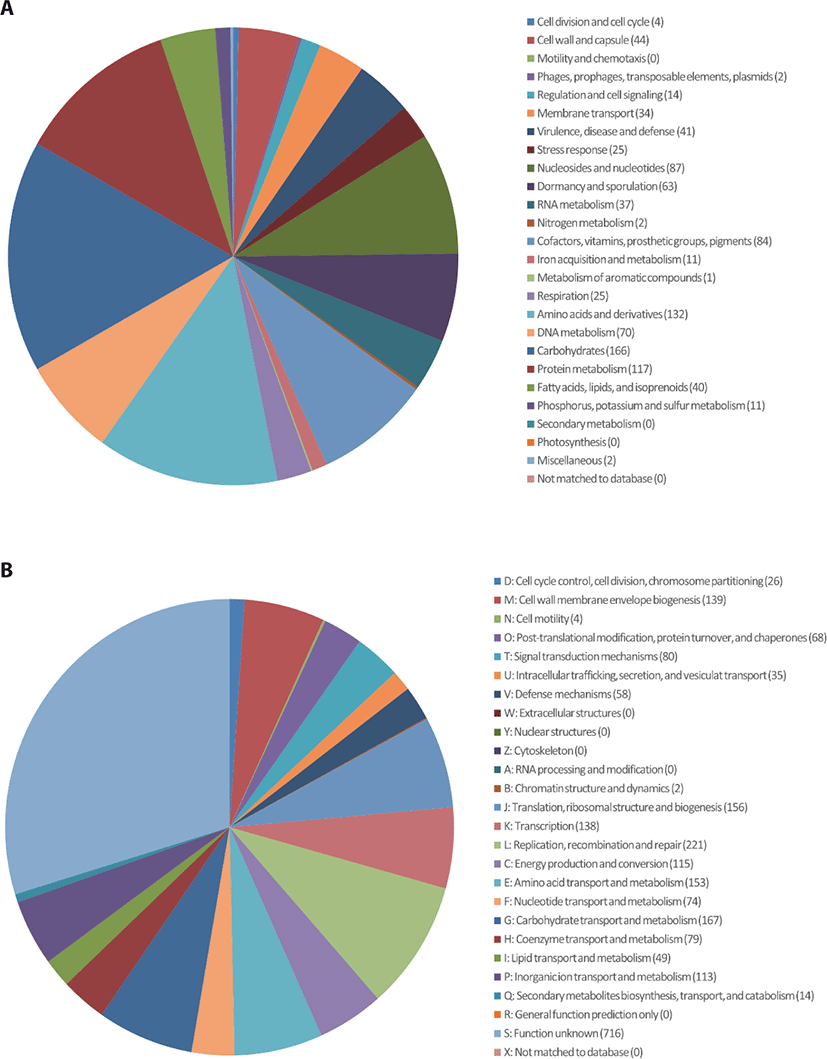
C. perfringens B20 was isolated from the feces of chickens maintained in a Chung-Ang University-affiliated farm (Anseong, Korea). Strain B20 was routinely cultured anaerobically in Brain-Heart Infusion (BD Bacto, Sparks, MD, USA) medium at 37°C for 24 h [5]. Genomic DNA was extracted from 12 h cultures using a QIAamp PowerFecal DNA Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The genomic DNA obtained was sequenced commercially at ChunLab (Korea) using the Pacific Biosciences (PacBio, Menlo Park, CA, USA) RSII Single Molecule Real-Time (SMRT) platform with a 20-kb SMRTbell™ template library. De novo assembly of the PacBio reads was performed using the PacBio SMAR Analysis ver. 2.3.0 program, and genome annotation was performed based on Rapid Annotation using Subsystem Technology (RAST) with default parameters (https://rast.nmpdr.org/) and CLgenomics™ ver. 1.55 software. Transfer RNAs (tRNAs) were identified using tRNAscan-SE ver. 1.3.1 [6], whereas non-coding ribosomal RNAs (rRNAs) were identified using INFERNAL ver. 1.1.3 software in conjunction with the Rfam 12.0 database [7]. The complete genome of C. perfringens B20 (Fig. 1) is 2,982,563 bp long with a G + C (guanine + cytosine) content of 28.2% and is assembled into two contigs with an N50 value of 2,925,822 bp. The genome contains 2,668 protein-coding sequences, 30 rRNA, and 94 tRNA genes (Table 1). Functional annotation of protein-coding sequences (Fig. 2) was performed using PRODIGAL ver. 2.6.2 software [8] and compared with the selected protein databases (SwissProt, KEGG, SEED, EggNOG).
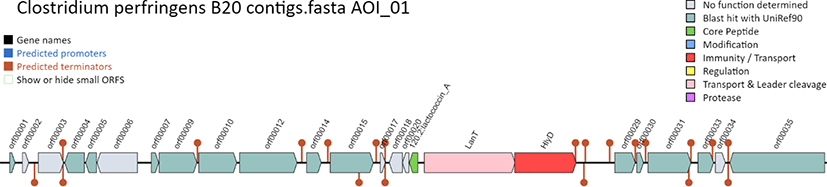
| Attribute | Value |
|---|---|
| Genome size (bp) | 2,982,563 |
| GC content (%) | 28.2 |
| No. of contigs | 2 |
| Total genes | 2,792 |
| Protein-coding gene | 2,668 |
| tRNA | 94 |
| rRNA | 30 |
| Plasmids | 0 |
| GenBank accession No. | JAGKIL000000000 |
Functional annotation revealed the presence of several multidrug resistance efflux pumps and tetracycline resistance genes. Putative bacteriocin-encoding gene clusters were determined in silico using the BAGEL4 online tool (http://bagel4.molgenrug.nl/) which revealed two areas of interest in contig 53CPB201_2, corresponding to the core bacteriocin, transport, and immunity genes (Fig. 3). One open reading frame encoding the core peptide of lactococcin A (E value, 4.37 × 10−49) was also detected. The amino acid sequence of the core peptide, deduced from the putative bacteriocin gene, showed a single peptide, non-pediocin-like bacteriocin, thereby indicating that it could be classified as a class IId bacteriocin [9]. Lactococcin A was initially isolated from Lactococcus lactis subsp. cremoris LMG 2130, in which it was located on a plasmid, and showed a narrow spectrum of activity against lactococci [10]. Additionally, LanT and HylD, encoding a bacteriocin ATP binding cassette (ABC) transporter and bacteriocin binding protein, respectively, were detected downstream of the core peptide gene, and they conceivably play functional roles in membrane-associated immunity. Moreover, a putative 63-amino acid-long bacteriocin, which was not identified using BAGEL4, was detected using RAST, with the following amino acid sequence: MENLNLNQLENINGGSTLGWATATATQLGLAAAATAEVPPLSAGLAAAGAV LAGGVAVYEAWH. A BLASTP search revealed identical protein sequences associated with C. perfringens, indicating that the putative bacteriocin gene could be novel. Genomic analyses provide evidence on bacteriocin production by C. perfringens B20 and could potentially enhance its pathogenicity by eliminating the Lactococcus community and other sensitive microorganisms in the chicken gut, thereby making disease control more challenging.