INTRODUCTION
Yeonsan ogye is a chicken breed native to Korea. The nutritional quality, taste, overall acceptance, and texture of the meat of the Korean native chicken breed, including the content of taste-related compounds, are superior to those of commercial broilers [1,2]; thus, consumers have a higher preference for it. Therefore, it would be a good strategy to crossbreed native chickens to produce broilers with tastier meat by taking advantage of the unique meat quality characteristics of native chickens. A study of the characteristics of the Yeonsan ogye chicken meat reported higher crude fat content and shearing power in Yeonsan ogye compared to other Korean native chickens, and the color of the meat was expressed as black [3]. In addition, it has been reported that the breast meat extract of Korean native chickens can alleviate inflammation [4].
To improve the palatability of chicken products, many studies have been conducted to determine the factors that influence meat quality. Studies have identified several parameters that influence meat quality, including tenderness, water-holding capacity, and meat color, in relation to the physical composition, while pH, dry matter, moisture, and intramuscular fat content are related to the chemical composition of the meat [5–8]. Taste-active precursors in meat, such as free amino acid (FAA) and nucleotide (NT), undergo chemical reactions during cooking, which determines many aspects of flavor, one of the main sensory features of meat [9–11].
As reported in [9], glycine, alanine, serine, threonine, and proline in the form of FAAs provide the sweet taste to the meat after cooking. Regarding NTs, inosine 5’-monophosphate, alone or in combination with other flavor compounds, provides an umami taste, greatly affecting the flavor of meat [12].
Many studies have been conducted to identify the genes related to the physicochemical properties of chickens. In [13], a study of the association of the fatty acid composition in chicken was conducted, and in [14], a gene related to the lipid profile of chicken muscle was reported. In [15,16], association studies were conducted on body composition. However, there have been no attempts to identify the genes related to FAA and NT content, which determine the flavor of chicken, in the whole genome. Furthermore, the traits present in the chicken Quantitative Trait Loci Database do not include the FAA and NT contents of breast meat [17]. Therefore, the objective of this study was to identify the potential candidate genes related to different FAA and NT contents in chicken breast through a whole-genome association analysis.
MATERIALS AND METHODS
For the construction of the experimental chicken population, a reciprocal crossbred F2 generation was produced using the Yeonsan ogye breed and the White Leghorn breed owned by the National Institute of Animal Science (NIAS, Korea). A total of 810 birds from the F2 generation were made up of two lines according to the sex combination of F0 birds and were raised until 10 weeks of age with the mixing of males and females. All F2 birds were raised on a farm at the NIAS Center for Livestock Genetic Resources under the same environmental and nutritional conditions. Overall experimental outline and the construction of the F2 generation is illustrated in Fig. 1.
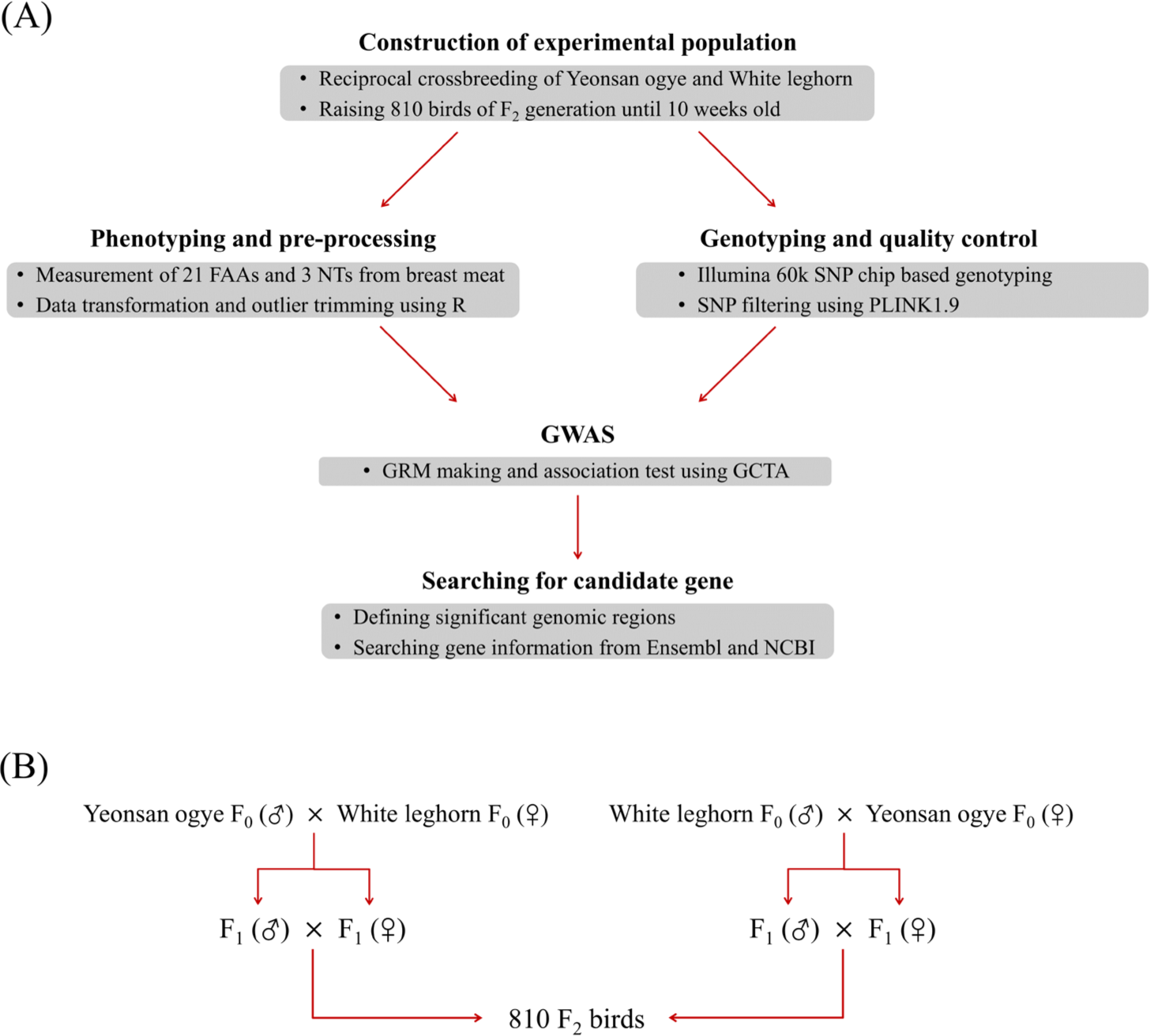
At 10 weeks of age, all birds were slaughtered on the same day at a commercial abattoir. Viscera were removed from the chicken carcass immediately after slaughter, rapidly frozen at −35°C, and then stored at −20°C at NIAS (Wanju, Korea) on the same day. Frozen samples were stored for 2–6 weeks, and then transported to Chungnam National University (Deajeon, Korea) and stored at −80°C until deboning. Next, samples were thawed at 4°C for 20 h, the breast meat was separated from the carcass, and then the breast meat was divided into samples for the FAA and NT component analyses. The FAA analysis was performed using a liquid chromatography system (UltiMate 3000RS, Thermo Fisher Scientific, Waltham, MA, USA) to quantify the content of 21 FAAs from a sample (Table 1). From the quantified FAA results, the essential FAAs and non-essential FAAs contents of chickens were added and each phenotype was used in an association analysis. For the NT content analysis, high-performance liquid chromatography (1200 Series, Agilent, Santa Clara, CA, USA) was applied. The resulting phenotypes were used in the association test as traits for the content of each substance and are summarized in Table 1.
CV, coefficient of variation; Min, minimum; Max, maximum; Trans, transformation; Sqrt, square root scaling; GABA, γ-aminobutyric acid; Min-Max, minimum-maximum scaling; Log, Log scaling; FAA, free amino acid content; AMP, adenosine monophosphate; NT, nucleotide content; IMP, inosine 5’-monophosphate.
To normalize the phenotypic data, a transformation was carried out to accurately estimate the effect of single-nucleotide polymorphisms (SNPs) in the association test. For each phenotype in the dataset, a Shapiro–Wilk normality test was performed using R software with the “shapiro.test()” function, and a transformation step was applied to the phenotype with a p-value of < 0.05. To find the optimal transformation method, the dataset with the highest p-value was adopted after performing normality tests on the raw data and data transformed by log, square root, square, and minimum/maximum (min/max) methods (Table 1). Then, to remove outliers, values of three standard deviations or more from the mean of each data point were treated as missing data, and all subjects whose analysis results were identified as missing data were excluded from the analysis. The FAA and NT component datasets were filtered in the quality control process, leaving 764 and 767 birds for the later analysis of FAA and NT components, respectively.
For genomic DNA (gDNA) extraction, blood collection was performed from the brachial veins of birds at 8 weeks of age. gDNA was extracted from the collected blood using the Wizard Genomic DNA Purification Kit (Promega, Madison, WI, USA), and the quality and concentration of the DNA were checked using a NanoDrop 1000 spectrophotometer (Thermo Fisher Scientific), with samples stored at −20°C. A total of 57,636 SNP genotypes were produced from the extracted gDNA using the Illumina 60K Chicken SNP BeadChip. The SNPs were removed if any of the following conditions were not met: call rate > 0.9, minor allele frequency > 0.01, Hardy–Weinberg equilibrium test p-value of > 0.01 × 10−6. After filtering, 29,175 SNPs were available for the subsequent analysis. PLINK 1.9 software was used for genotype quality control procedure [18].
A genetic relationship matrix (GRM) was calculated using the genome-wide complex trait analysis (GCTA) tool[19]. An association analysis was performed using GCTA’s mixed linear model leave-one-chromosome-out statistical method, and the following model was used:
where, y is a vector of the phenotype value, including FAAs and NTs; a is the mean value of phenotypic record; b is the additive effect of the SNP to be tested for association; x is the genotype of the SNP to be tested; g− is the effect of all SNPs captured by the GRM, except those on the chromosome where SNP testing is located; and e is the residual effect vector. To accurately estimate the marker effect, sex, experimental batch, half-sibling family, and body weight were used as covariates. After the association analysis, the significance of the SNP effect was evaluated by a Bonferroni corrected p-value of 5%: p < 0.05/number of SNPs use for the association test. The GWAS result was visualized with a Manhattan plot using R software.
Based on the chicken reference genome (GRCg6a), we assumed that the genomic region around 1 Mbp was the significant region centered on the position of the most significant SNP for each chromosome, and known genes included in that region were reported as candidate genes. Location-based gene information was obtained by searching the Ensembl (https://asia.ensembl.org) and NCBI (https://www.ncbi.nlm.nih.gov) databases.
RESULTS
The descriptive statistics of the phenotype measurements of FAAs and NTs from the F2 chicken population are presented in Table 1. For these phenotypic values, basic statistical analyses were performed after transformation and outlier trimming. Histograms and quantile–quantile plots are shown in Fig. 2. Lysine, for which most values were measured as zero, was excluded from the test because its distribution was judged to be inappropriate for the association test. It was determined that the cause of this inappropriate data was an error in phenotype measurement.
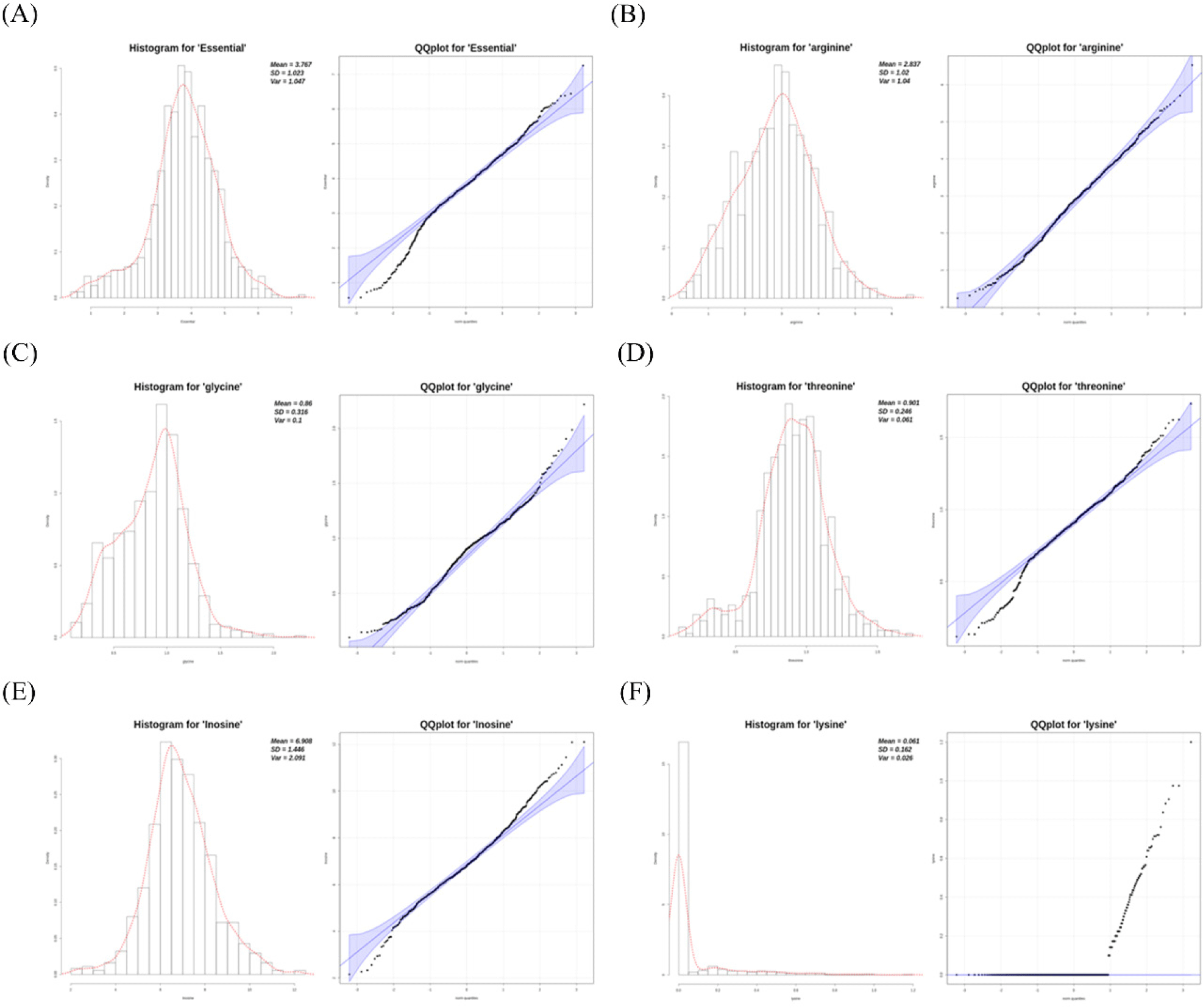
Table 2 shows the significant SNPs for each trait as a result of the association test, with results presented as summary statistics of the GWAS. The SNP markers were located at chicken (Gallus gallus) chromosome 1 (GGA1), GGA2, GGA3, GGA4, and GGA7. Of the three SNPs associated with the essential FAAs, Gga_rs15943371 and Gga_rs14162297 were located at bp 33,096,495 and 33,432,575 on GGA2, respectively, and Gga_rs16609168 was located at bp 31,241,647 on GGA7. In arginine, Gga_rs14162297 did not reach the significance level, but it was the second most significant SNP among the markers located on GGA2, and Gga_rs16609168 was also the SNP with the highest significance in GGA7. Another SNP related to arginine was Gga_rs13971906, which was located at bp 170,582,107 on GGA1, and was significant for inosine. As other SNPs associated with inosine, GGaluGA265714, GGaluGA265712, and GGaluGA265969 were located on GGA4 at bp 74,314,272, 74,285,145, and 75,827,200, respectively. The SNP associated with glycine was GGaluGA028030 and was located at bp 81,831,988 on GGA1. The SNP associated with threonine was Gga_rs14412837 and was located at bp 108,537,571 on GGA3.
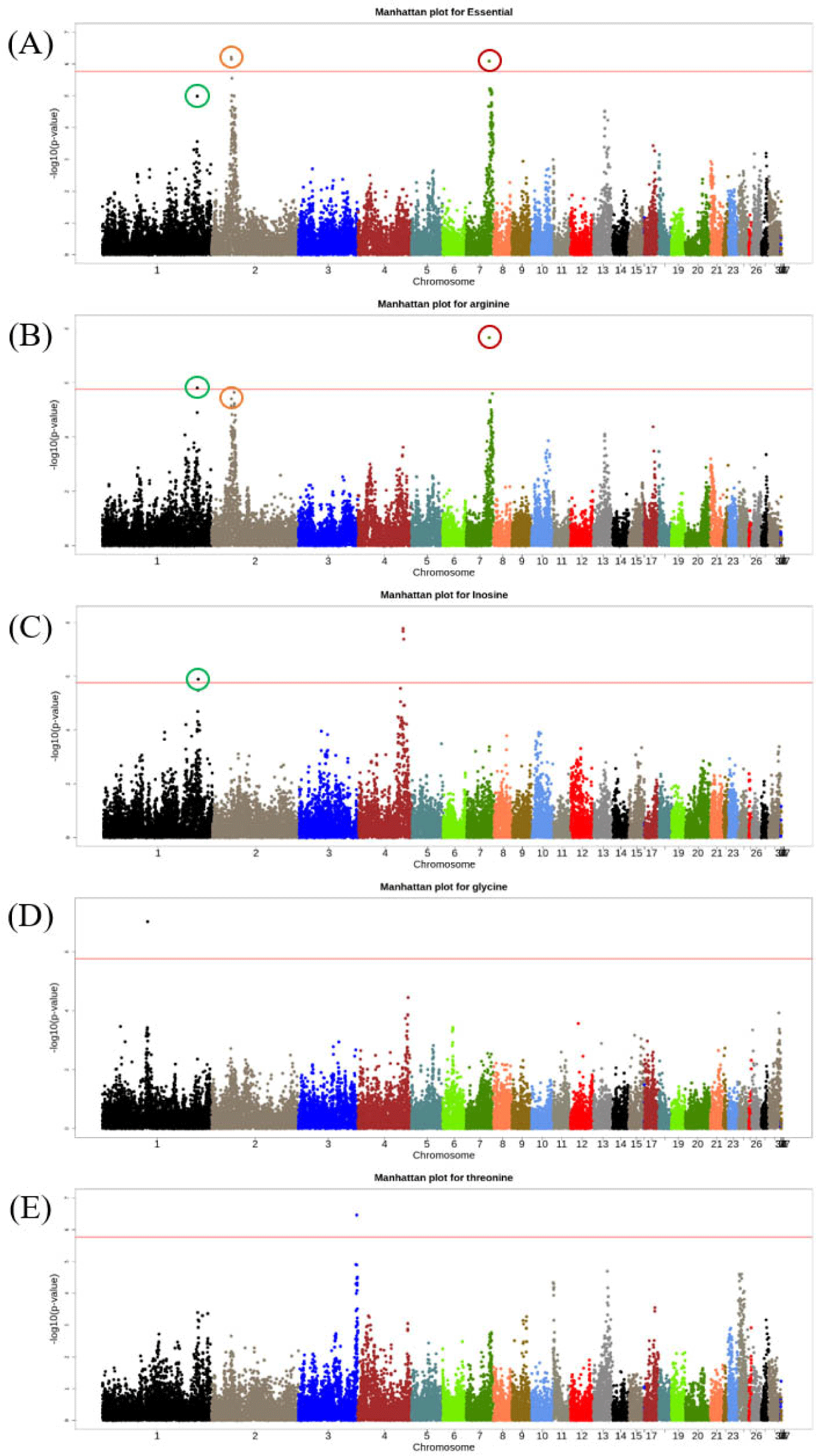
The Manhattan plot in Fig. 3 presents only the traits with significant SNPs. The most significant SNP on GGA1 was rs13971906 for the essential FAAs, arginine, and inosine. Gga_rs14162297, which had the second highest significance for the essential FAAs, also had the second highest significance for arginine. Table 3 shows the significantly related genomic regions for each trait and the genes included therein.
DISCUSSION
In this study, the candidate genes related to FAA and NT contents of chicken breast meat were identified through the GWAS analysis. In the GWAS result, the trend of significance for the essential FAAs was similar to that of arginine (Fig. 3). The reason why the two traits shared a significant genomic region in GGA1, GGA2, and GGA7 was that arginine content accounted for the largest portion of the essential FAAs in the phenotype. The arginine content in the breast meat of the F2 crossbred chicken was greater than the content of all other FAAs when the mean values were compared.
From the GWAS results of inosine and arginine, it was observed that the genomic region on GGA1 was commonly significantly related (Table 2). One of the genes included in this region, the KCNRG gene, encoded a voltage-gated potassium channel regulator protein. The voltage-gated potassium channel can regulate the nitric oxide synthesis pathway, which is a process in which L-arginine is converted to nitric oxide. This might have an effect on the arginine content in the body [20,21]. In addition, the voltage-gated potassium channel can modulate the insulin sensitivity of human skeletal muscle, which modulates the nitric oxide pathway and affects L-arginine transport and metabolism. Therefore, the potassium channel may be related to the amount of L-arginine in muscle [22–24]. Interestingly, if the content of adenosine triphosphate (ATP), a glucose metabolite in the muscle, changes as insulin manages skeletal muscle glucose uptake, the content of inosine, which is a repeatedly degraded form of ATP, may also be affected [25,26]. For the same reason, it could be concluded that the KCNIP4 gene on the significant region in GGA4 encodes a potassium channel interacting protein.
Diet and nutritional level in birds are factors affecting the chemical composition of muscles. Among the candidate genes related to dry matter intake in cattle reported in Javaid et al. [27] and Brunes et al. [28], the RB1, LPAR6, RCBTB2, CYSLTR2, FNDC3A, MLNR, PHF11, and KPNA3 genes were identical to the genes in the genomic region commonly related to inosine and arginine in this study. Therefore, it is possible that the difference in nutritional condition according to the feed intake of chickens raised under ad libitum conditions in this study affected the chemical composition of breast meat [27].
It has been reported that the arginine content can be increased by reducing nitric oxide synthase when the expression of the HOXA3 gene among the GGA2 genes related to arginine content in Table 3 is regulated [29]. It has also been confirmed that mutations in the THSD7B gene of GGA7 affect nitric oxide metabolism [30]. In the GWAS result of the threonine content, the MMUT gene encoding the methylmalonyl-CoA mutase located on GGA3 was included as a candidate gene. In animals, including chickens, threonine is catabolized to alpha-ketobutyrate by serine/threonine dehydratase, then converted to propionyl-CoA and carboxylated to methylmalonyl-CoA [31–33]. The catalyzed product is finally converted to succinyl-CoA by methylmalonyl-CoA mutase and used in the tricarboxylic acid cycle [34]. Therefore, threonine content may be affected by methylmalonyl-CoA mutase involved pathway which use threonine as a substrate.
The molecules with significantly related SNP markers in this study were arginine, inosine, threonine, and glycine. As taste-active precursors that determine meat flavor during cooking, arginine and inosine give a bitter taste, and threonine and glycine give a sweet taste [9]. This study had limited success in finding a significant gene region related to the contents of FAA and NT molecules that give umami and sour tastes in chicken breast. The reason that significant SNPs were not found through GWAS in most traits is thought to be due to the large impact of environmental effects on the FAA and NT contents. The FAA content increases as protease hydrolyzes meat protein, and this degradation proceeds even in frozen meat [35]. A significant change in the amino acid profile of meat according to the storage duration at freezing temperature has also been reported [36]. In addition, in the process of thawing frozen meat, several NT-metabolizing enzymes perform a chain degradation to change the NT content [37–39]. Based on the results of this study, it could be concluded that the period of storage under freezing conditions before analysis after the slaughter of chickens was long enough to result in significant changes in the FAA and NT contents. Future studies may capture more genetic effects when association tests are conducted by performing phenotyping on meat with a shorter storage time than in the present study.
CONCLUSION
Changes in the concentrations of FAAs and NTs in chicken are the main factors that change the flavor. To our knowledge, this was the first GWAS to explore genetic markers related to the content of FAAs and NTs in chicken. Through the GWAS, nine significant SNPs related to essential FAAs, arginine, glycine, threonine, and inosine components were found, and six significant genomic regions and candidate genes included therein were reported. These genomic regions were mapped on chromosome 1, 4, and 7 of chicken, and the potential candidate genes were KCNRG for arginine and inosine, HOXA3 and THSD7B for arginine, KCNIP4 for inosine, and MMUT for threonine. The reported candidate genes might be important genetic markers in chicken selection and breeding to help meet consumer expectations for healthier, tastier, and flavored meat.
















