INTRODUCTION
An elite calf refers to a calf with superior abilities in economic aspects such as growth and reproduction. Among them, for steers, those with superior meat yield grade and meat quality grade related to carcass traits are highly useful as beef cattle. For cows, they are highly useful as breeding cattle that steadily produce calf through pregnancy and childbirth, and contributes greatly to the farm income. Production of elite calves is achieved through systematic and continuous selection and improvement. Improvements in economic traits have been made through selection [1], and for selection, as the process of selecting parents to pass on superior genetic abilities to the future generation, it is important to estimate the abilities of the parents through genetic evaluation. There are two ways of selection for the Hanwoo (Korea cattle) improvement industry. From bulls that were raised with the same breeding management, the performance test selects candidate bulls through growth report verification while the progeny test selects a Korean proven bull’s number (KPN) through verifying the growth and carcass grade of the candidate bull’s progeny [2]. The selected KPN supplies semen to farms across the country to lead the improvement of Hanwoo but selecting less than 30 bulls per year for new KPN selection to improve Hanwoo reduces the genetic diversity of Hanwoo and farmer’s preference for specific KPN has become a factor for the rise of the inbreeding coefficient. Therefore, the decrease in genetic diversity is accelerating [3]. Even if the same KPN is used, there is a large variation in the carcass grade of the offspring. This is because the genetic ability of the cow, which is responsible for 50% of the calf’s genetic ability, is not considered at all. It has been reported that selection based on KPNs lacks efficiency due to limitations in time, cost and selection intensity [4]. In order to solve the genetic diversity and dependence on specific KPN, a breeding plan that takes into account the genetic ability of cows is necessary, and to estimate the genetic ability of cows, the pedigree and phenotype information of individuals is necessary. For cows, estimation is very difficult because the pedigree and ability test report (estimated breeding value [EBV], performance, pedigree eta.) are insufficient in the farms. This has led to the early culling of cows with superior genetic resources [5]. However, due to the animal products traceability (APT) system implemented in 2010, phenotype and pedigree registration information collection became possible by individual identification number search through the Korea Institute for Animal Products Quality Evaluation (KAPE) and Korea Animal Improvement Association (KAIA) and this has laid the foundation for genetic evaluation. If this is used for elite cow selection and Hanwoo improvement, the problem of inbreeding can be solved through securing a large number of genetic diversity and it can increase the profitability of farms and have improvement effect as the reproduction through elite cows becomes possible, instead of elite calf production through specific KPN. Therefore, this study analyzed the EBV and accuracy using the pedigree information of Hanwoo cows in Gyeongnam to construct basic data for the selection of elite cows.
MATERIALS AND METHODS
For the test group, 34,705 individual identification numbers of Korean cattle born in the Gyeongnam collected from 2003 to 2017 year was provided by GAST, the university enterprise of Gyeongsang National University. The sex, region, birth year and pedigree were collected by searching the individual identification number through the KAPE and KAIA. From the searched information, individuals who did not have a pedigree or had a discrepancy in the gender, region, individual identification number or pedigree were considered as outliers and removed. Ultimately, 33,128 cows were used for analysis. And shown Table 1. Pedigree were constructed by tracing the individual identification number of the test group back to four generations through the KAIA. For ease of analysis, it was sorted by Animal, Sire, and Dam, and the pedigree viewer version 4.0 program (Brian Kinghorn, University of New England, Armidale, Australia) was used for the pedigree structure and renumbering. Afterward, the R program (http://cran.r-project.org) suitable for large-scale information processing was used to search and delete duplicate objects and correct discrepancies in individual identification numbers. Finally, pedigree for 48,358 heads were constructed. Out of these, 674 heads were represented as KPNs and 37,838 heads were represented as cows. There were 2 heads with unknown KPNs and 232 heads with unknown cows. And 13,456 heads with unknown KPNs and cows, it excluded from the analysis.
In order to analyze the test group’s EBV, a reference group is required, along with pedigree and phenotype. The pedigree estimates the relationship coefficient between individuals using the pedigree relationship of the test and reference groups, and it is analyzed with the phenotype to estimate the EBV. The reference group used for the analysis was 545,483 heads slaughtered at an average age of 30 months and it was provided by the KAPE. The pedigree collection was done in the same way as the previous test group’s pedigree construction method, and a total of 1,270,300 heads pedigree were constructed. For the phenotype, the carcass trait was measured after 24 hours of refrigeration after the slaughter in accordance with the Livestock Grade Determination Standard Detail #2014-4 posted by the Ministry of Agriculture, Food and Rural Affairs (MAFRA). The carcass weight (CWT) was measured as the sum of the left and right frozen body weights, and the eye muscle area (EMA) was measured by incising between the left and right thoracic vertebrae and the first lumbar spine at a right angle to the vertebrae and using an area ruler to find the area of the last thoracic vertebrae. The backfat thickness (BFT) was measured along the right side of the EMA where it entered 2/3 of the way in at the abdomen and the marbling score (MS) was measured with the naked eye by comparing the degree of fat deposition in the muscle, located at the EMA measurement site, to the reference table (1 = devoid and 9 = abundant).
The pedigree of the test and the reference groups were combined to remove duplicate individuals using the R program, and ultimately, pedigree for 1,309,511 heads were constructed and used for analysis. Pedigree and phenotype were used to estimate the EBV, prediction error variance (PEV), genetic parameters, and genetic correlation for each trait and the multiple trait animal model of the BLUPF90 program [6] were used. For the fixed effect, the birth year, birth month and age at slaughter were used, and the mixed model equation is as follows [7].
Here, Y is the vector for the observed value of economic traits, X is the vector for the fixed effect of birth year, birth month, and age at slaughter, and the vector for the estimated value of the fixed effects. Z is the vector for the random effects of individuals and u is the vector for the estimated value of individuals and e is the vector for random errors. E(y) = Xβ, Var(u) = G = A , Cov(µ,e) = 0 is assumed to give Var(y) = V = ZGZ′ + R. Here, A is the numeric relationship matrix (NMR) constructed by the pedigree between individuals and is the additive genetic variance and is the random environmental variance.
The accuracy of EBV was estimated using PEV and additive genetic variance by trait from the solution values of the BLUPF90 program analysis result and the equation is as follows.
Here, Acc is the accuracy of the EBV, PEV is the predicted error variance of the EBV, and is the additive genetic variance.
RESULTS AND DISCUSSION
The reference group used in this study were 545,483 heads that were slaughtered at an average age of 30 months. The mean and standard deviation for CWT, EMA, BFT, and MS were 431.7 ± 43 kg, 91.1 ± 9.7 cm2, 13.2 ± 4.5 mm, and 5.7 ± 1.9 points, respectively, and they are shown in Table 2.
| Trait | Mean | SD | Min. | Max. |
|---|---|---|---|---|
| CWT (kg) | 431.7 | 43 | 303 | 551 |
| EMA (cm2) | 91.1 | 9.7 | 65 | 116 |
| BFT (mm) | 13.2 | 4.5 | 1 | 25 |
| MS (point) | 5.7 | 1.9 | 1 | 9 |
Looking at the measured carcass of the proved cattle for the national genetic evaluation, the CWT, EMA, BFT, and MS were reported by Hwang et al. [8] as 321.01 ± 41.89 kg, 75.72 ± 8.19 cm2, 8.27 ± 3.69 mm, and 2.91 ± 1.63 points, respectively and Lee et al. [9] reported them as 352.99 ± 37.83 kg, 82.35 ± 8.30 cm2, 7.99 ± 3.08 mm, and 3.05 ± 1.47 points, respectively. If you look at the carcass measured in commercial farms, Won et al. [10] reported by collecting steer from commercial farms in one rural Gangwondo region that the values of CWT, EMA, BFT, and MS were 426.8 ± 49.3 kg, 87.9 ± 9.1 cm2, 10.9 ± 4.1 mm, and 5.6 ± 2.0 points, respectively. Dang et al. [11] reported from the steer of the Pyeongchang region that the values of CWT, EMA, BFT, and MS were 423.91 ± 44.90 kg, 91 ± 10.11 cm2, 13.63 ± 5.33 mm, and 5.6 ± 1.83 points, respectively. The proved cattle are slaughtered at 24 months of age to be used in the national genetic evaluation. However, since traits such as intramuscular fat grow rapidly after 24 months when the skeletal and muscle growth of the livestock is complete, there is a difference in the carcass as shipment happens at 29 to 31 months, which is the peak in growth. It is considered that the accuracy of EBV is rather low due to the use of data measured in the early and middle stages of growth for traits that develop in the later stages of growth. Therefore, it is considered that a higher accuracy estimation could be possible by establishing and utilizing a reference group that has a similar carcass as the Korean cattle group shipped nationwide. The carcass of the reference group used in this study are similar to those of the previous studies [10,11] and is considered to be a suitable group for the genetic evaluation conducted for commercial farms.
Henderson proposed the BLUP that analyzes the EBV by constructing an NRM from common ancestors using pedigrees between individuals within a group, and this method requires pedigree, which is the connection point between the individual and the group. Hanwoo registration information is divided into pre-registration, basic registration, pedigree registration, and advanced registration. Pre-registration refers to domestically produced cattle that are recognized as Hanwoo in terms of appearance, and basic registration refers to cattle that are 6 months of age or older that only know their own information with 70 points for cows and 75 points for steers and bulls. In addition, pedigree registration refers to cattle whose parents are above the basic registration level and those that do not have disqualification conditions in appearance. Advanced registration refers to cattle within 24 months and 36 months of age that have pedigree registration and have passed the appearance and ability test [12]. Based on the paternity confirmation project of the APT system implemented in 2010, the change of pedigree structure of the test group and the subsequent EBV change were analyzed and the results are shown in Table 3 and Fig. 1.
| Birth year | Basic | Pedigree | Advanced | Total | |||
|---|---|---|---|---|---|---|---|
| Head | Percent | Head | Percent | Head | Percent | ||
| 2003–2009 | 603 | 25 | 562 | 24 | 1,206 | 51 | 2,371 |
| 2010–2017 | 3,194 | 10 | 20,847 | 68 | 6,716 | 22 | 30,757 |
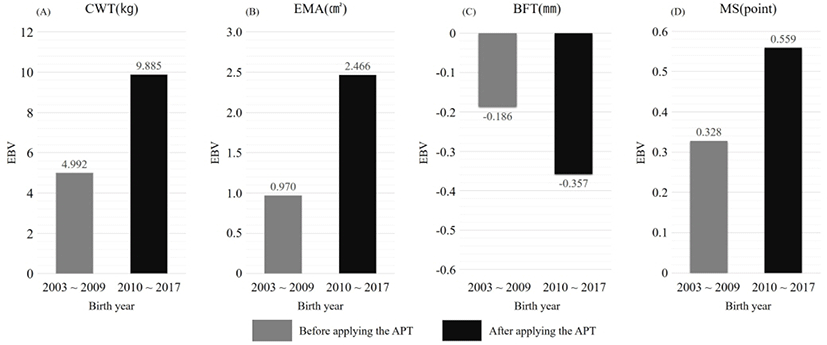
Looking at Table 3, there were a total of 2,371 cows, born between 2003 to 2009, from which 603 cows had basic registration (25%), 562 cows had pedigree registration (24%) and 1,206 cows had advanced registration (51%). The proportion of pedigree registered cows was relatively low but it gradually increased and reached a point of 20,847 cows (68%) between 2010 to 2017. The proportion of registered cows (pedigree registration, advanced registration) with accurate parental information was 90% (pedigree registration cows: 68%, advanced registration cows: 22%), which was a 15% increase compared to 2003 to 2009. It can be seen that 90%, excluding basic registration cows, have pedigree. Looking at Fig. 1, the EBV was 4.992 kg, 0.970 cm2, −0.186 mm, and 0.328 points in the order of CWT, EMA, BFT, and MS for the cow group born between 2003 to 2009. It can be confirmed that the EBV of the cow group born between 2010 to 2017 improved to 9.885 kg, 2.466 cm2, −0.357 mm, and 0.559 points. In order to analyze the EBV, the proportion of registered cows (pedigree registration, advanced registration) with accurate parental information is important. Due to the paternity confirmation project of the APT system implemented in 2010 and the change in awareness of pedigree registration importance in the farms, the proportion of registered cows (pedigree registration, advanced registration) with accurate parental information have greatly increased, which shows that the pedigree accuracy of farms has increased. It is considered that through this, the high accuracy of genetic ability results have increased the improvement efficiency of Hanwoo and therefore increased the EBV.
The generic variance, residual variance, heritability and accuracy results estimated in this study are shown in Table 4. The heritability of the CWT, EMA, BFT, and MS estimated in this study were 0.587, 0.416, 0.476, and 0.571. Looking at previous studies, Utrera and Van Vleck [13] surveyed 72 papers published between 1962 to 2004 and reported the average heritability of carcass traits, in the order of CWT, EMA, BFT, and MS, to be 0.40, 0.40, 0.36 and 0.37. Roh et al. [14] reported 0.28, 0.35, 0.39, and 0.51, and Sun et al. [15] reported 0.39, 0.30, 0.33, and 0.62. Do et al. [16] reported 0.28, 0.23, 0.20, and 0.28, and Shin et al. [17] reported 0.360, 0.442, 0.479, and 0.581. Most of the previous studies showed a middle and high degree of heritability and were consistent with the results of this study.
| Trait | Additive variance | Residual variance | Heritability | Accuracy |
|---|---|---|---|---|
| CWT | 1,178.65 | 828.124 | 0.587 | 0.559 |
| EMA | 40.848 | 57.256 | 0.416 | 0.551 |
| BFT | 10.467 | 11.513 | 0.476 | 0.554 |
| MS | 2.208 | 1.658 | 0.571 | 0.558 |
The degree of accuracy of CWT, EMA, BFT, and MS estimated in this study were 0.559, 0.551, 0.554, and 0.558, respectively. Looking at previous studies, Lee et al. [9] reported 0.6047, 0.6748, 0.6748, and 0.7364 as the degree of accuracy for CWT, EMA, BFT, and MS for the test group of the progeny test cattle, which are higher values than this study. Shin et al. [17] reported 0.441, 0.458, 0.464, and 0.472 as the degree of accuracy for CWT, EMA, BFT, and MS for commercial farms, which are lower values than this study. Since the progeny test cattle have a high pedigree relationship within the group, a high accuracy estimation is possible despite the small reference group size. But it is reported that commercial farms have low connection points in the pedigree of the test group and reference group, resulting in the difficulty in estimating accuracy [18].
The heritability of the trait and the accuracy of the EBV are strongly influenced by the genetic relations within the reference group and the genetic variation of the trait [18]. Pszczola et al. [19] reported that more diverse genetic variations can be used when the average pedigree relationship between the individuals forming the reference group is lower, and more accurate relationship coefficient can be estimated for the test group to be selected when there is a higher pedigree relationship with the reference group, leading to increased values or more accuracy. Therefore, it can be seen that the EBV and the numerical value of the accuracy to be analyzed are greatly influenced by the size of the reference group and the connection point between the groups. In the case of genetic evaluation for farms, as the size of the reference group increases, the success rate of genetic evaluation of various individuals increases because various pedigree relationships can be estimated. In the farms, the connection points with the reference group increase with higher pedigree relationship of an individual, which enables accurate relationship coefficient analysis. Through this, it is seen that the heritability and accuracy were estimated higher than previous studies because of the lowered residual variance of each trait and high relationship coefficient for the EBV and the degree of accuracy analyzed in this study.
The genetic and phenotype correlations estimated in this study are shown in Table 5. The genetic correlation was within the −0.024 to 0.576 range, and the genetic correlation of the EMA and BFT was −0.024. All traits except for those with negative correlation showed to have a positive correlation. The highest value was 0.576 for the genetic correlation between the CWT and the EMA, and the lowest value was −0.024 for the genetic correlation between the EMA and the BFT. On the other hand, the phenotype correlation between traits ranged from 0.037 to 0.483, showing a positive correlation in all traits. Looking at previous studies, the genetic correlation between CWT and EMA was reported by Roh et al. [14], Hwang et al. [8], Do et al. [16] as 0.651, 0.63, and 0.80, respectively and Veseth et al. [20] that studied the foreign breed Hereford reported 0.58. The genetic correlation between CWT and BFT was reported by Dang et al. [11] and Do et al. [16] as 0.36 and 0.17, respectively. The genetic correlation between CWT and MS was reported by Park et al. [21] and Do et al. [16] as 0.20 and 0.21. The genetic correlation between EMA and BFT was reported by Yoon et al. [22], Roh et al. [14], and Salces et al. [23] as −0.21, −0.139, and −0.17, respectively. The genetic correlation between EMA and MS was reported by Salces et al. [23] and Roh et al. [24] as 0.12 and 0.401, respectively. The genetic correlation of BFT and MS was reported by Hwang et al. [8] and Park et al. [21] as 0.04 and −0.02, respectively. Most previous studies showed similar correlations with the results of this study, and as suggested by Kim et al. [25], it was similar to the direction of Hanwoo improvement at the national level. According to a study by Lande [26], genetic correlation is defined as a biological mechanism consisting of the pleiotropy and linkage disequilibrium of genes. It reported that improvement in the desired direction can be made if the selection uses individuals with functional alleles that apply commonly to both of those traits.
| Trait | CWT | EMA | BFT | MS |
|---|---|---|---|---|
| CWT | 0.576 | 0.355 | 0.152 | |
| EMA | 0.483 | −0.024 | 0.469 | |
| BFT | 0.333 | 0.037 | 0.029 | |
| MS | 0.141 | 0.389 | 0.067 |
CONCLUSION
Since 2010, pedigree has been systematically managed through the paternity confirmation project of Hanwoo following the implementation of the APT system, and it significantly increased the proportion of Hanwoo cows in the Gyeongnam region with pedigree registration. It can be confirmed that, excluding basic registration cows, 90% of the cows have accurate parental information. Through pedigree with higher reliability, genetic evaluation of farms became possible. Based on this, the improvement efficiency increased through selection by farm and as a result, the EBV increased. It is considered that in the future, maintaining reliable pedigree through steady paternity confirmation project participation by farms and building a reference group of various size and diverse genetic composition will be important factors in Hanwoo improvement. Through this, if elite cow groups are formed and selected, the possibility of elite calf reproduction through elite cows by farms will increase, rather than elite calf reproduction that is dependent on specific KPN. The effect of cow improvement and a decrease in the specific KPN preference phenomenon will increase the profitability of farms.

